Book assembled from https://www.assembla.com/spaces/silk/wiki/ on 07-Feb-2013 by vladimir.alexiev@ontotext.com
About Silk
The Silk Link DIscovery Framework is a tool for discovering relationships between data items within different Linked Data sources. Data publishers can use Silk to set RDF links from their data sources to other data sources on the Web.
The official homepage of Silk can be found on http://www4.wiwiss.fu-berlin.de/bizer/silk/.
Silk Workbench
Introduction
Silk Workbench is a web application which guides the user through the process of literlinking different data sources.
The Silk Workbench offers the following features:
- It enables the user to manage different sets of data sources and linking tasks.
- It offers a graphical editor which enables the user to easily create and edit link specifications.
- As finding a good linking heuristics is usually an iterative process, the Silk Workbench makes it possible for the user to quickly evaluate the links which are generated by the current link specification.
- It allows the user to create and edit a set of reference links used to evaluate the current link specification.
The Silk Workbench provides the following components:
- Workspace Brower Enables the user to browse the projects in the workspace. Linking Tasks can be loaded from a project and committed back to it later.
- Linkage Rule Editor A graphical editor which enables the user to easily create and edit link specifications. The widget will show the current link specification in a tree view while allowing editing using drag-and-drop.
- Evaluation Allows the user to execute the current Link Specification. The links are displayed while they are generated on-the-fly. Generated links for which the reference link set does no specify their correctness, the user may confirm or decline their correctness. The user may request detailed summaries on how the similarity score of specific links is composed of.
The typical workflow of creating a new Link Specification consists of:
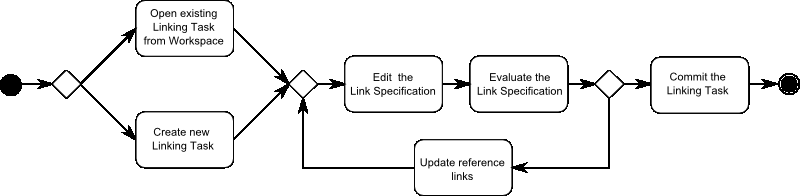
- The user opens an existing Linking Task from the Workspace or creates a new Linking Task.
- The user uses the Linkage Rule Editor to refine the current Linkage Rule.
- The output of the Link Specification is evaluated based on the reference links using the Evaluation.
- If all links are correct, the user commits the Link Specification to the Workspace. If some links are wrong the user proceeds with the next step.
- The user confirms or declines the correctness of a number of links.
Workspace
The workspace allows users to manage data sources and linking tasks defined for each project. The Workspace Browser shows a tree vies of all Projects in the current Workspace:
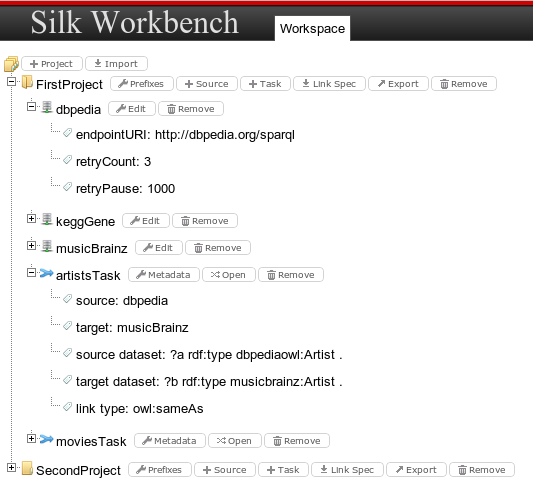
Projects
A project holds the following information:
- All URI prefixes which are used in the project.
- A list of data sources.
- A list of linking tasks.
Users are able to create new projects or import existing ones. Existing projects can be deleted or exported to a single file.
Data sources
A data sources holds all information that is needed by Silk to retrieve entities from it. Users can add new data sources and edit their properties:
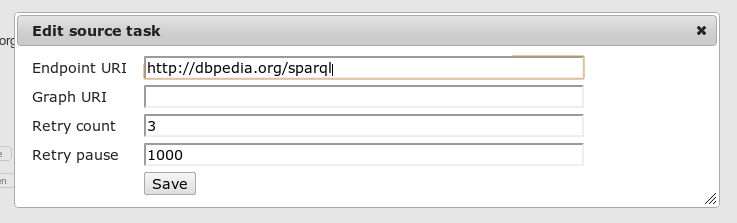
The following properties can be edited:
- Endpoint URI The URI of the SPARQL endpoint
- Graph URI Only retrieve instances from a specific graph
- Retry count To recover from intermittent SPARQL endpoint connection failures, the ‘retryCount’ parameter specified the number of times to retry connecting.
- Retry pause To recover from intermittent SPARQL endpoint connection failures, the ‘retryPause’ parameter specifies how long to wait between retries.
Linking Tasks
A linking tasks consists of the following elements:
- Metadata
- A link specification
- Positive and negative reference links
Linking Tasks can be added to a existing project and removed from it. Clicking on Metadata opens a dialog to edit the meta data of a linking task:
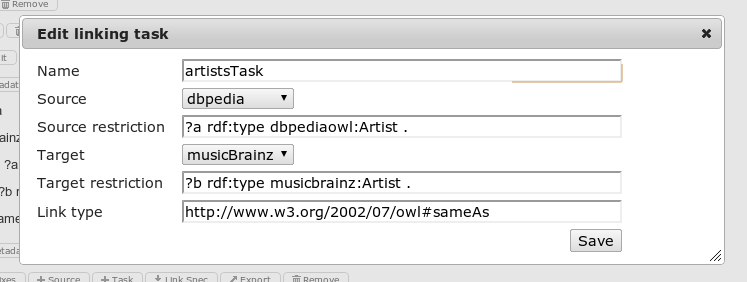
The following properties can be edited:
- Name The unique name of the linking task
- Source The source data set
- Source restriction Restricts source dataset using SPARQL clauses
- Target The target data set
- Target restriction Restricts target dataset using SPARQL clauses
- Link type Type of the generated link e.g. owl:sameAs
Clicking on the open button opens the Link Specification Editor
Linkage Rule Editor
The linkage Rule Editor allows users to edit linkage rules in graphical way. Linkage rules are created as a tree, resembling the Silk LSL, by dragging and dropping the rule elements.
The editor is divided in two parts:
- The left pane contains the most frequent used property paths for the given data sets and restrictions. It also contains a list of all Silk operators (transformations, comparators and aggregators) as draggable elements.
- The right part (editor pane) allows for drawing the flow chart by combining the elements chosen.
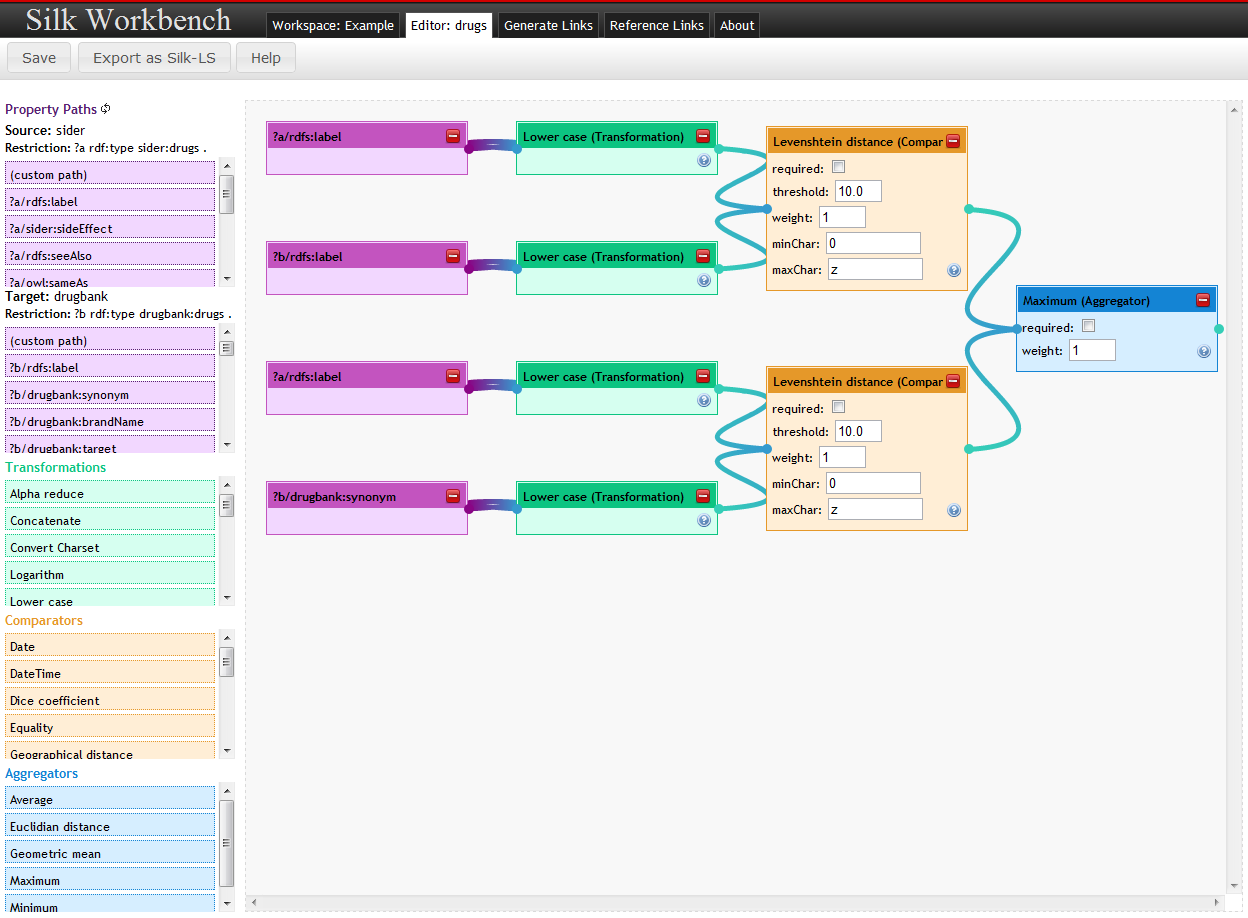
Editing
- Drag elements from the left pane to the editor pane
- Connect the elements by drawing connections from and to the element endpoints (dots to the left and right of the element box)
- Build a flow chart by connecting the elements, ending in one single element (either a comparison or aggregation)
The editor will guide the user in building the flow chart by highlighting connectable elements when drawing a new connection line.
Property Paths
Property paths for both data sources to be interlinked are loaded on the left pane and added in the order of their frequency in the data source.
Users can also add custon paths by dragging the (custom path) element to the editor pane and editing the path.
Operators
The following operator panes are shown below the property paths:
Hovering over the operator elements will show you more information on them.
Threshold
Threshold defines the minimum similarity of two data items which is required to generate a link between them. Please provide values between 0 and 1.
Link Limit
Link Limit defines the number of links originating from a single data item. Please choose between 1 and n (unlimited).
Generating Links
As finding a good linking heuristics is usually an iterative process, the Silk Workbench allows the user to quickly evaluate the links which are generated by the current link specification.
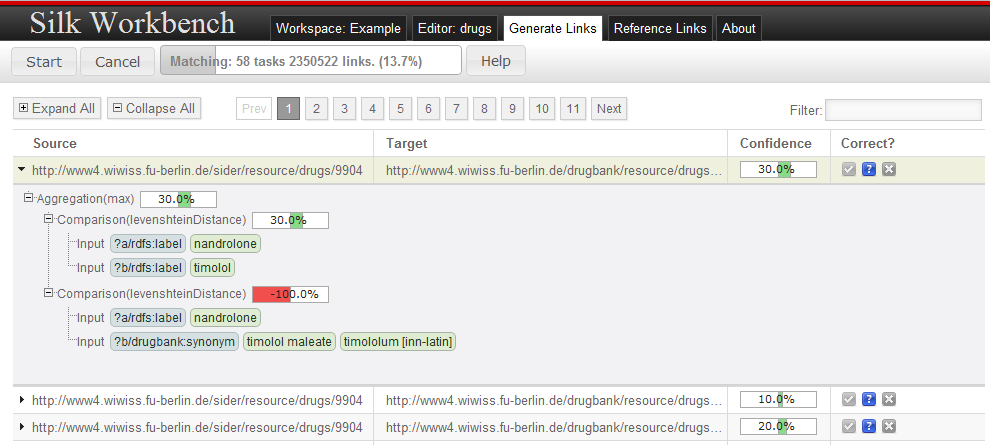
After clicking on the Start button, the linking engine starts to generate links in the background. The view is updated whenever new links have been found to show the all generated links. For further examination, a drill-down view can be shown by clicking on a link.
The drill-down shows a detailed summary how the individual comparisons and aggregations contribute to the overall fitness of the link, the overall similarity between two links is composed by clicking on a link. This information allows the user to spot parts of the similarity evaluation which did not behave as expected.
Based on its correctness, each link can be associated to one of the following 3 categories:
 Confirms the link as correct. Confirmed links are part of the positive reference link set.
Confirms the link as correct. Confirmed links are part of the positive reference link set. Contains link whose correctness is not decided i.e. which are not contained in the reference link sets.
Contains link whose correctness is not decided i.e. which are not contained in the reference link sets. Confirms the link as incorrect. Incorrect links are part of the negative reference link set.
Confirms the link as incorrect. Incorrect links are part of the negative reference link set.
In order to evaluate the correctness of a link specification, the user typically wants to evaluate the correctness of links which are close the similarity threshold. Clinking on the confidence header sorts all links by their confidence which allows to find these links. In addition, a filter can be added, so that only links are shown which contain a specific string.
Managing Reference Links
In order to iteratively increase the quality of a link specification, the workbench holds a set of reference links for which their correctness has been confirmed or declined by the user.
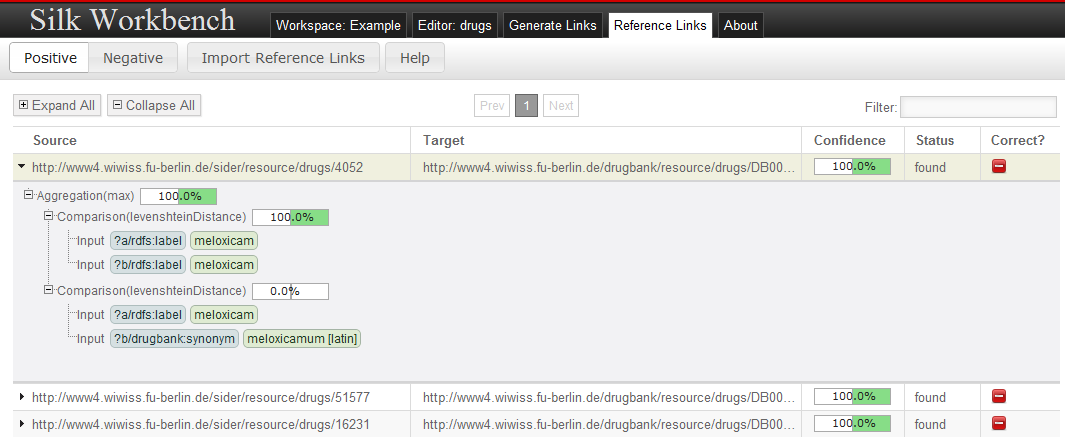
Reference Links may be imported and exported in the Ontology Alignment Format specified at http://alignapi.gforge.inria.fr/format.html.
Learning Linkage Rules
The Silk Workbench supports learning linkage rules. In order to be able to learn linkage rules, the current linking task must contain reference links.
Usually reference links can be added in two ways:
- By loading existing reference links from a file. See Managing Reference Links for details.
- By using the sampling tab
After reference links have been added, the learning can be started by pressing the start button. While learning, the current population of candidate linkage rules is displayed. At any time, the learning can be stopped by pressing the stop button. The learning will stop automatically as soon as either the full f-Measure is reached or the maximum number of iterations has been exceeded.
As soon as the learning has finished, the user can view the learned linkage rules and select a linkage rule for loading it into the editor.
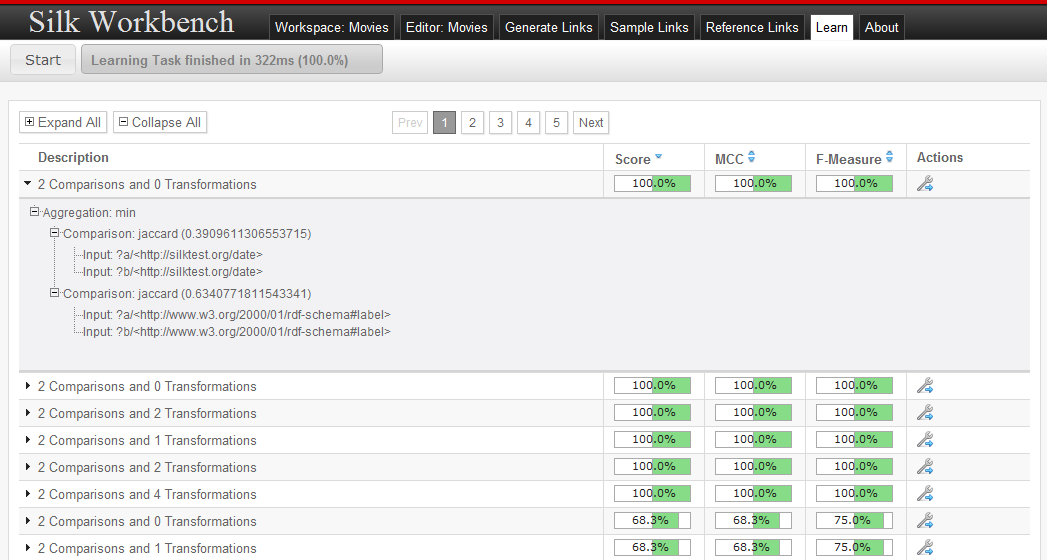
Basic Concepts
This section introduces the basic concepts in link discovery such as data sources, linkage rules and reference alignments.
Data Sources
Overview
Data sources hold the access parameters to local or remote SPARQL endpoints or RDF files. The defined data sources may later be referred to and used by their ID. Data Sources can by defined using either the API or XML.
Available Data Sources
SPARQL Endpoint Data Source Definitions
For SPARQL endpoints (dataSource type: sparqlEndpoint) the following parameters exist:
| Parameter | Description | Default |
|---|---|---|
| endpointURI | The URI of the SPARQL endpoint | |
| login | Login required for authentication | No login |
| password | Password required for authentication | No password |
| instanceList | a list of instances to be retrieved. If not given, all instances will be retrieved. Multiple instances can be separated by a space. | Retrieve all instances |
| pageSize | Limits each SPARQL query to a fixed amount of results. Silk implements a paging mechanism which translates the pagesize parameter into SPARQL LIMIT and OFFSET clauses. | 1000 |
| graph | Only retrieve instances from a specific graph. | |
| pauseTime | To allow rate-limiting of queries to public SPARQL servers, the pauseTime statement specifies the number of milliseconds to wait in between subsequent queries. | 0 |
| retryCount | To recover from intermittent SPARQL endpoint connection failures, the retryCount parameter specifies the number of times to retry connecting. | 3 |
| retryPause | Specified how long to wait between retries. | 1000 |
Example (XML)
<DataSOurce id="dbpedia" type="sparqlEndpoint">
<Param name="endpointURI" value="http://dbpedia.org/sparql" />
<Param name="retryCount" value="100" />
</DataSOurce>
Example (Scala API)
Note that all parameters except the endpoint URI optional and can be left out.
Source("dbpedia",
SparqlDataSource(
endcpointURI = "http://dbpedia.org/sparql",
login = "user",
password = "password",
graph = "http://dbpedia.org",
pageSize = 1000,
pauseTime = 0,
retryCount = 3,
retryPause = 1000
)
)
RDF File Data Source Definitions
For RDF files (dataSource type: file) the following parameters exsit:
| Parameter | Description | Default |
|---|---|---|
| filr (mandatory) | The location of the RDF file. | |
| format(mandatory) | The format of the RDF file. Allowed values: “RDF/XML”, “N-TRIPLE”, “TURTLE”, “TTL”, “N3” |
Currently the data set is held in memory.
Example (XML)
<DataSource id="musicbrainz" type="file">
<Param name="file" value="musicbrainz_dump.nt" />
<Param name="format" value="N-TRIPLE" />
</DataSource>
Example (Scala API)
Source("musicbrainz",
FileDataSource(
file = "musicbrainz_dump.nt",
format = "N-TRIPLE"
)
)
Linkage Rule
A linkage rule specifies how two data items are compared for similarity. A linkage rule consists of four basic components:
- Path Input Retrieves values from a entity by a given RDF path e.g.
?movie/dbpedia:director/rdfs:label - Transformation Applies a data transformation to all values e.g.
lowerCase - Comparison Evaluates the similarity of two inputs based on a user-defined distance measure and returns a confidence.
- Aggregation Aggregates multiple confidence values.
Path Input
Overview
An input retrieves all values which are connected to the entities by a specific path.
Every path statement begins with a variable (as defined in the datasets), which may be followed by a series of path elements. If a path cannot be resolved due to a missing property or a too restrictive filter, an empty result set is returned.
The following operators can be used to traverse the graph:
| Operator | Name | Use | Description |
|---|---|---|---|
| / | forward operator | <path_segment>/<property> |
Moves forward from a subject resource (set) through a property to its object resource (set). |
| \ | reverse operator | <path_segment>\<property> |
Moves backward from an object resource (set) through a property to its subject resource (set). |
| [ ] | filter operator | <path_segment>[<property> <comp_operator> <value>] <path_segment>[@lang <comp_operaor> <value>] |
Reduces the currently selected set of resources to the ones matching the filter expression. comp_operator may be one of >, <, >=, <=, =, != |
Example (XML)
# Select the English label of a movie
<Input path="?movie/rdfs:label[@lang='en']" />
# Select the label (set) of the director(s) of a movie
<Input path="?movie/dbpedia:director/rdfs:label" />
# Select the albums of a gicen artist (albums have an dbpedia:artist property)
<Input path="?artist\dbpedia:artist[rdf:type=dbpedia:Album]" />
Example (Scala API)
# Select the English label of a movie
Path.parse("?movie/rdfs:label[@lang='en']")
# Select the label (set) of the director(s) of a movie
Path.parse("?movie/dbpedia:director/rdfs:label")
# Select the albums of a given artist (albums have an dbpedia artists propetry)
Path.parse("?artist\dbpediaLartist[rdf:type=dbpedia:Album]")
Transformation
Overview
As different datasets usually use different data formats, a transformation can be used to normalize the values prior to comparison.
Parameters
TODO
Example (XML)
<TransformInput function="lowerCase">
<TransformInput function="replcae">
<Input path="?a/rdfs:label?" />
<Param name="search" value="_" />
<Param name="replace" value=" " />
</TransformInput>
</TransformInput>
Example (Scala API)
TransformInput(
id = "ReplaceUnderscores",
transformet = ReplaceTransformer("_", " ")
inputs = PathInput(path=Path.parse("?a/rdfs:label"))
)
Transformations
Silk provides the following transformation and normalization functions:
| Function and parameters | Description |
|---|---|
| removeBlanks | Remove whitespace from a string. |
| removeSpecialChars | Remove special characters (including punctuation) from a string. |
| lowerCase | Convert a string to lower case. |
| upperCase | Convert a string to upper case. |
| capitalize(allWords) | Capitalizes the string i.e. converts the first character to upper case. If ‘allWords’ is set to true, all words are capitalized and no only the first character. By default ‘allWords’ is set to false. |
| stem | Apply word stemming to the sting. |
| alphaReduce | Strip all non-alphabetic characters from a string |
| numReduce | Strip all non-numeric characters from a string. |
| replace(string search, string replace) | Replace all occurrences of “search” with “replace” in a string. |
| regexReplace(string regex, string replace) | Replace all occurrences of a regex “regex” with “replace” in a string. |
| stripPrefix | Strip the prefix from a string. |
| stripPostfix | Strip the postfix from a string. |
| stipUriPrefix | Strip the URI prefix (e.g. http://dbpedia.org/resource/) from a string. |
| concat | Concatenates strings grom two inputs. |
| logarithm([base]) | Transforms all numbers by applying the logarithm function. Non-numeric values are left unchanged. If base is no defined, it defaults to 10. |
| convert(string sourceCharset, string targetCharset) | Converts the string from “sourceCharset” to “targetCharset” |
| tokenize([regex]) | Splits the string into tokens. Splits at all matches of “regex” if provided and at whitespaces otherwise. |
| removeValues(blacklist) | Removes specific values (i.e. stop words) from the value set. ‘blacklist’ is a comma-separated list of words. |
Comparison
Overview
A comparison operator evaluates two inputs and computes the similarity based on a user-defined distance measure and a user-defined threshold.
The distance measure always outputs 0 for a perfect match, and a higher value for an imperfect match. Only distance values between 0 and threshold will result in a positive similarity score. Therefore it is important to know how the distance measures work and what the range of their output values is in order to set a threshold values sensibly.
Parameters
| Parameter | Description |
|---|---|
| required (optional) | If required is true, the parent aggregation only yields a confidence value if the given inputs have values for both instances. |
| weight (optional) | Weight of this comparison. The weight is used by some aggregations such as the weighted average aggregation. |
| Hreshold | The maximum distance. For normalized distance measured, the threshold should be between 0.0 and 1.0. |
| distanceMeasure | The used distance measure. For a list of available distance measures see below. |
| Inputs | The 2 inputs for the comparison. |
Example (XML)
<Compare metric="leenshteinDistance" threshold="2.0" required="true">
<TransformInput function="lowerCase">
<Input path="?a/rdfs:label" />
</TransformInput>
<TransformInput function="lowerCase">
<Input path="?b/rdfs:label" />
</TransformInput>
</Compare>
Example (Scala API)
Comparison(
id = "labels",
required = false,
weight = 1,
threshold = 2.0,
metric = LevenshteinDistance()
inputs = PathInput(path = Path.parse("?a/rdfs:label")) ::
PathInput(path = Path.parse("?b/rdfs:label")) :: Nil
)
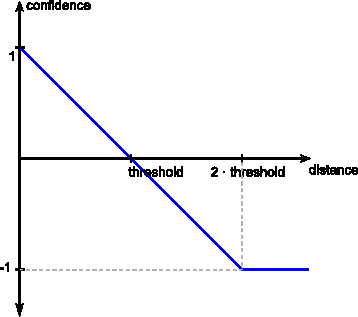
Threshold
The threshold is used to convert the computed distance to a confidence between -1.0 and 1.0. Links will be generated for confidences above 0 while higher confidence values imply a higher similarity between the compared entities.
Distance Measures
Character-Based Distance Measures
Character-based distance measures compare strings on the character level. They are well suited for handling typographical errors.
| Measure | Description | Normalized |
|---|---|---|
| levenshteinDistance | Levenshtein distance. The minimum number of edits needed to transform on string into the other, with the allowable edit operations being insertion, deletion, or substitution of a single character | No |
| levenshtein | The levenshtein distance normalized to the interval [0, 1] | Yes |
| jaro | Jaro distance metric. Simple distance metric originally developed to compare person names | Yes |
| jaroWinkler | Jaro-Winkler distance measure. The Jaro-Winkler distance metric is designed and best suited for short strings such as person names | Yes |
| equality | 0 if strings are equal, 1 otherwise | Yes |
| inequality | 1 if strings are equal, 0 otherwise | Yes |
Example (XML)
<Compare metric="levenshteinDistance" threshold="2">
<Input path="?a/rdfs:label" />
<Input path="?b/gn:name" />
</Compare>
Token-Based Distance Measures
While character-based distance measure work well for typographical errors, they are number of tasks where token-base distance measures are better suited:
- Strings where parts are reordered e.g. “John Doe” and “Doe, John”
- Texts consisting of multiple words
| Measure | Description | Normalized |
|---|---|---|
| jaccard | Jaccard distance coefficient | Yes |
| dice | Dice distance coefficient | Yes |
| softjaccard | Soft Jaccard similarity coefficient. Same as Jaccard distance but values within an levenshtein distance of ‘maxDistance’ are considered equivalent. | Yes |
Example (XML)
<Compare metric="jaccard" threshold="0.2">
<TransformInput function="tokenize">
<Input path="?a/rdfs:label" />
</TransformInput>
<TransformInput function="tokenize">
<Input path="?b/gn:name" />
</TransformInput>
</Compare>
Special Purpose Distance Measures
Silk offers a number of distance measures which are designed to compare specific types of data e.g. numeric values.
| Measure | Description | Normalized |
|---|---|---|
| num(float minValue, float maxValue) | Computes the numeric difference between two numbers. Parameters: minValue, maxValue The minimum and maximum values which occur in the datasource. |
No |
| date | Computes the distance between two dates (“YYYY-MM-DD” format). Returns the difference in days. | No |
| dateTime | Computes the distance between two date time values (xsd:dataTime format). Returns the difference in seconds. | No |
| wgs84(string unit, string curveStyle) | Computes the geographical distance between two points. Parameters: unit The unit in which the distance is measured. Allowed values: “meter” or “m” (default), “kilometer” or “km” |
No |
Example (XML)
<Compare metric="wgs84" threshold="50">
<Input path="?a/wgs:84:geometry" />
<Input path="?b/wgs84:geometry" />
<Param name="unit" value="km" />
</Compare>
Aggregation
Overview
An aggregation combines multiple confidence values into a single value. In order to determine if two entities are duplicates it is usually not sufficient to compare a single property. For instance, when comparing geographic entities, and aggregation may aggregate the similarities between the names of the entities and the similarities based on the distance between the entities.
Parameters
Required (Optional)
The required attribute can be set if the aggregation only should generate a result if a specific suboperator return a value
Weights (Optional)
Some comparison operators might be more relevant for the correct establishment of a link between two resources than others. For example, depending on data formats/quality, matching labels might be considered less important than matching geocoordinates when linking cities. If this modifier is not supplied, a default weight of 1 will be assumed. The weight is only considered in the aggregation types average, quadraticMean and geometricMean.
Type
The function according to the similarity values are aggregated. The following functions are included in Silk:
| Id | Name | Description |
|---|---|---|
| average | AverageAggregator | Evaluate to the (weighted) average of confidence values. |
| max | MaximumAggregator | Evaluate to the highest confidence in the group |
| min | MinimumAggregator | Evaluate to the lowest confidence in the group |
| quadraticMean | QuadraticMeanAggregator | Apply Euclidian distance aggregation |
| geometricMean | GeometricMeanAggregator | Compute the (weighted) geometric mean of a group of confidence values |
Example (XML)
<Aggregate type="average">
<Compare metric="jaro" required="true">
<Input path="?a/rdfs:label" />
<Input path="?b/gn:name" />
</Compare>
<Compare metric="num">
<Input path="?a/dbpedia:populationTotal" />
<Input path="?b/gn:population" />
</Compare>
</Aggregate>
Example (Scala)
Aggregation(
id = "id1",
required = false,
weight = 1,
operators = operators,
aggregator = MaximumAggregator()
)
Output
Overview
An output represents a destination where the generated links are written to. Outputs can have an acceptance windows (defined by minConfidence and maxConfidence) e.g. for separating accepted links and links with a lower confidence which need to be verified before being accepted.
Example (XML)
<Outputs>
<Output type="output type" minConfidence="lower threshold" maxConfidence="upper threshold">
...
</Output>
</Outputs>
Example (Scala API)
Output(
id = "MyOutput",
writer = FileWriter(file = "output.nt", format = "ntriples")
)
Available Output Types
File Output
| Parameter | Description |
|---|---|
| file | Writes the links to a file. Links are written to {user.dir}/.silk/output by default. |
| format | The output format. Available formats are “ntriples” and “alignment” |
Example (XML)
<Outputs>
<Output type="file" minConfidence="0.1">
<Param name="file" value="accept_links.nt" />
<Param name="format" value="ntriples" />
</Output>
</Outputs>
Formats
N-Triples
Writes the links as N-Triples statements.
Alignment
Writes the links in the OAEI Alignment Format. This includes not only the uris of the source and target entities, but also the confidence of each link.
SPARQL/Update Output
| Parameter | Description |
|---|---|
| uri | The URI of the SPARQL/Update endpoint e.g. http://localhost:8090/virtuoso/sparql |
| login | Login required for authentication |
| password | Password required for authentication |
| parameter | The HTTP parameter used to submit queries. Defaults to “query” which works for most endpoints. Some endpoints require different parameters e.g. Sesame expects “update” and Joseki expects “request”. |
| graphUri | The URI of the graph to put the links |
Example (XML)
<Outputs>
<Output type="sparul">
<Param name="uri" value="http://localhost:8080/query" />
</Output>
</Outputs>
Detailed Alignment (Work in Progress)
Writes the links in a detailed alignment format.
Example (XML)
<DetailedAlignment>
<Cell>
<Entity1 rdf:resource="http://dbpedia.org/resource/Hydroflumenthiazide"/>
<Entity2 rdf:resource="http://www4.wiwiss.fu-berlin.de/drugbank/resource/drugs/DB00774" />
<Aggregate similarity="1.0">
<Compare similarity="1.0">
<Input path="?a/rdfs:label">
<Value>Idroflumetiazide</Value>
<Value>Hydroflumethiazide</Value>
</Input>
<Input path="?b/rdfs:label">
<Value>Hydroflumethiazide</Value>
</Input>
</Compare>
<Compare similarity="1.0">
<Input path="?a/rdfs:label">
<Value>Idroflumetiazide</Value>
<Value>Hydroflumethiazide</Value>
</Input>
<Input path="?b/drugbank:synonym">
<value>Idroflumetiazide</value>
</Input>
</Compare>
</Aggregate>
</Cell>
...
</DetailedAlignment>
Alternative (as RDF/XML)
<?xml version='1.0' encoding='utf-8' standalone='no'?>
<rdf:RDF xmlns:rdf='http://www.w3.org/1999/02/22-rdf-syntax-ns#'>
<DetailedAlignment>
<map>
<Cell>
<entity1 rdf:resource="http://dbpedia.org/resource/Hydroflumethiazide"/>
<entity2 rdf:resource="http://www4.wiwiss.fu-berlin.de/drugbank/resource/drugs/DB00774"/>
<aggregate>
<similiarity>1.0</similarity>
<compare>
<similiarity>1.0</similarity>
<input>
<path>?a/rdfs:label</path>
<value>Idroflumetiazide</value>
<value>Hydroflumethiazide</value>
</input>
<input>
<path>?b/rdfs:label</path>
<value>Hydroflumethiazide</value>
</input>
</compare>
<compare>
<similiarity>1.0</similarity>
<input>
<path>?a/rdfs:label</path>
<value>Idroflumetiazide</value>
<value>Hydroflumethiazide</value>
</input>
<input>
<path>?b/drugbank:synonym</path
<value>Idroflumetiazide</value>
</input>
</compare>
</aggregate>
</Cell>
</map>
...
</DetailedAlignment>
</rdf:RDF>
Reference Links
Overview
Reference Links are a set of links whose correctness has either been confirmed or declined by the user. Reference links can be used to evaluate the completeness and correctness of a linkage rule.
We distinguish between positive and negative reference links:
- Positive reference links represent definitive matches
- Negative reference links represent definitive non-matches
Link Specification Language (LSL)
The Silk framework provides a declarative language for specifying which types of RDF links should be discovered between data sources as well as which conditions data items must fulfill in order to be interlinked. This section describes the language constructs of the Silk Link Specification Language (Silk-LSL)
The example below gives an overview of the main language constructs of Silk-LSL
<Silk>
<Prefixes>
<Prefix id="rdfs" namespace="http://www.w3.org/2000/01/rdf-schema#" />
<Prefix id="dbpedia" namespace="http://dbpedia.org/ontology/" />
<Prefix id="gn" namespace="http://www.geonames.org/ontology#" />
</Prefixes>
<Datasources>
<Datasource id="dbpedia">
<Param name="endpointURI" value="http://demo_sparql_sever1/sparql" />
<Param name="graph" value="http://dbpedia.org" />
</Datasource>
<Datasource id="geonames">
<Param name="endpointURI" value="http://demo_sparql_server2/sparql" />
<Param name="graph" value="http://sws.geonames.org/" />
</Datasource>
</Datasources>
[<Blocking blocks="100" />]
<Interlinks>
<Interlink id="cities">
<LinkType>owl:sameAs</LinkType>
<SourceDataset dataSource="dbpedia" var="a">
<RestrictTo>
?a rdf:type dbpedia:City
</RestrictTo>
</SourceDataset>
<TargetDataset dataSource="geonames" var="b">
<RestrictTo>
?b rdf:type gn:P
</RestrictTo>
</TargetDataset>
<LinkageRule>
<Aggregate type="average">
<Compare metric="jaro">
<Input path="?a/rdfs:label" />
<Input path="?b/gn:name" />
</Compare>
<Compare metric="num">
<Input path="?a/dbpedia:populationaTotal" />
<Input path="?b/gn:population" />
</Compare>
</Aggregate>
</LinkageRule>
<Filter threshold="0.9" />
<Outputs>
<Output type="file" minConfidence="0.95">
<Param name="file" value="accepted_links.nt" />
<Param name="format" value="ntriples" />
</Output>
<Output type="file" maxConfidence="0.95">
<Param name="file" value="verify_links.nt" />
<Param name="format" value="alignment" />
</Output>
</Outputs>
</Interlink>
</Interlinks>
</Silk>
Structure and Elements
The Silk-LSL is expressed in XML as specified by the corresponding Silk XML Schema. The root tag name is <Silk>. A valid document may contain four types of top-level statements beneath the root element:
- prefix definitions
- datasource definitios
- link specifications
- output definitions
<?xml version="1.0" encoding="utf-8" ?>
<Silk>
<Prefixes ... />
...
<Datasources ... />
...
[<Blocking ... />]
...
<Interlinks ... />
...
[<Outputs ... />]
...
</Silk>
The Blocking and Outputs statements are optional.
Prefix Definitions
Prefix definitions are top-level statements that allow the binding of a prefix to a namespace:
<Prefixes>
<Prefix id="prefix id" namespace="namespace URI" />
</Prefixes>
Example (XML)
<Prefixes>
<Prefix id="rdf" namespace="http://www.w3.org/1999/02/22-rdf-syntax-ns#" />
</Prefixes>
Data Source Definitions
Data source definitions are top-level statements that allow the specification of access parameters to local or remote SPARQL endpoints. The defined data sources may later be referred to and used by their ID within link specification statements.
<DataSources>
<DataSource id="data source ID" type="dataSource type">
<Param name="parameter name" value="parameter value" />
</DataSource>
</DataSources>
For details see Data Sources
Blocking Data Items
Since comparing every source resource to every single target resource results in a number of n * m comparisons (n being the number of source resources, m the number of target resources) which might be too time consuming, blocking can be used to reduce the number of comparisons. Blocking partitions similar data items into clusters reducing the comparisons to items in the same cluster.
For example given two datasets describing books, in order to reduce the number of comparisons, we could block the books by publisher. In this case only books from the same publisher will be compared. Given a number of 40,000 books in the first dataset and 30,000 in the second dataset, evaluating the full Cartesian product requires 1.2 billion comparisons.
If we block this datasets by publisher, each book will be allocated to a block based on its publisher. Using 100 blocks, if the books are uniformly distributed, there will be 300 respectively 300 books per block, which reduces the number of comparisons to 12 million.